Quantitative biology
Quantitative biology
Quantitative approaches to biology are taking an increasingly important role in understanding biological phenomena. The great advances in systems biology have shown that computational and mathematical approaches can lead to insights that are complementary to the more traditional approach. Bioinformatics tools are now commonplace in the context of genomics, molecular biology, and structural biology to better analyze biological data. In this spirit, our current work takes ideas from non-equilibrium statistical mechanics in physics and applies them to genetic evolution. Non-equilibrium physics is still an actively researched field with many open problems. However, some standard core techniques that lay at the heart of the theory are still unutilized in the context of evolutionary biology.
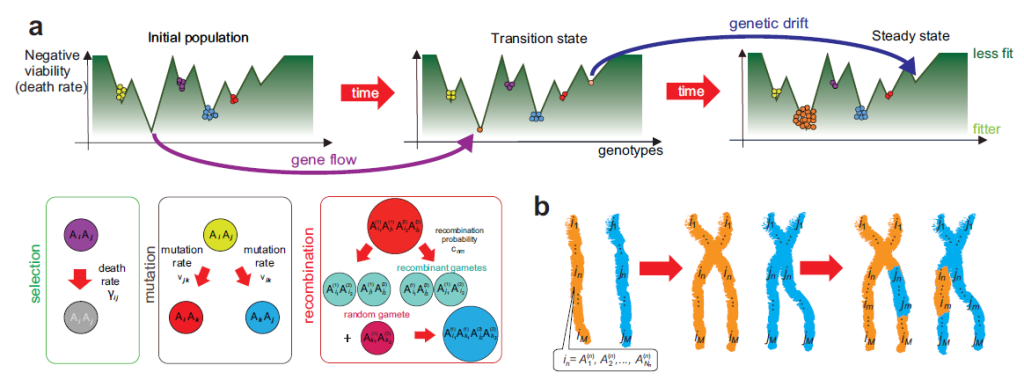
Fig 1: (a) Schematic of the evolution process for multiple loci evolving in time. (b) Our labeling for the recombination process.
We previously derived a master equation (this is standard physics nomenclature), which fully takes into account the fundamental processes that contribute to evolution: selection, mutation, recombination, gene flow, and genetic drift. This, in principle, can be applied to any evolving population of multicellular eukaryotic organisms limited to the assumptions laid out by the Hardy-Weinberg theory. In contrast to the standard Hardy-Weinberg equilibrium, we extend our scope to explicitly include a more realistic approach (e.g. the presence of selection, mutation, recombination, genetic drift, and gene flow in a non-equilibrium framework). Using simple input parameters such as the death rate of individuals and mutation and recombination rates, one can quantitatively predict the population distribution of various genotypes. We identify some interesting situations – such as a critical wait time before gene flow occurs – and derive a simple formula for them based on the master equation. The speed of propagation of a trait through filial generations can also be determined. Moreover, the viability of individual alleles can be assessed thereby giving way to foreshadowing the possible array of available traits in a population at any given time in the future. NYU Shanghai has some new labs to perform some of these ideas experimentally, we are currently in progress of doing that. We believe that our work is an important step towards quantitatively predicting the evolutionary process thus allowing techniques developed in physics to be brought into biology.